Find Care
Find a Location
All Location Types
All Specialties
Why Choose Houston Methodist?
Houston Methodist is one of the nation's leading health care systems and academic centers, providing unparalleled quality — and safety — in clinical care, advanced technology and patient experience. That is our promise of leading medicine.
Full Spectrum of Services
Leading Care Everywhere
Get Quality Care Now
Personalized Care
Full Spectrum of Services

Full Spectrum of Services
From annual checkups and mammograms to innovative,
customized treatment plans for complex conditions, we
offer a full spectrum of quality care for all your health care
needs.
Leading Care Everywhere
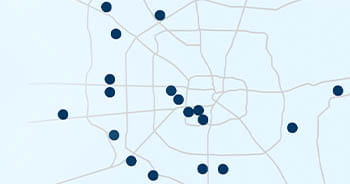
Leading Care Everywhere
Whether you need primary, specialty, emergency or hospital
care, Houston Methodist’s expertise is conveniently located
wherever you may need care.
Get Quality Care Now

Get Quality Care Now
Whether you’re suffering from allergies, a fever, a skin
irritation or a broken bone, Houston Methodist offers a
variety of ways to get the care you need when you need it.
Personalized Care

Personalized Care
We understand how important it is to treat you as an
individual, with you own unique health history and needs.
Your health care should be personalized, and we are
dedicated to establishing lifelong relationships, providing a
consistent care team and building customized plans
designed for your well-being.

Research & Innovation
At Houston Methodist, fostering innovations with the potential for clinical application is at the very heart of what we do — allowing us to bring cutting-edge treatments to patients faster. Our deep commitment to finding new ways to prevent and treat illness to improve the health of our patients means that our primary goal is to make a clinical trial available to every person who needs and wants to participate in one.
News & Health Information
On Health Blog — Lifestyle & Wellness
In The News — Recent Press
Events — Virtual & In-Person
Leading Medicine Blog — For Physicians
On Health Blog — Lifestyle & Wellness
In The News — Recent Press
Events — Virtual & In-Person
Leading Medicine Blog — For Physicians
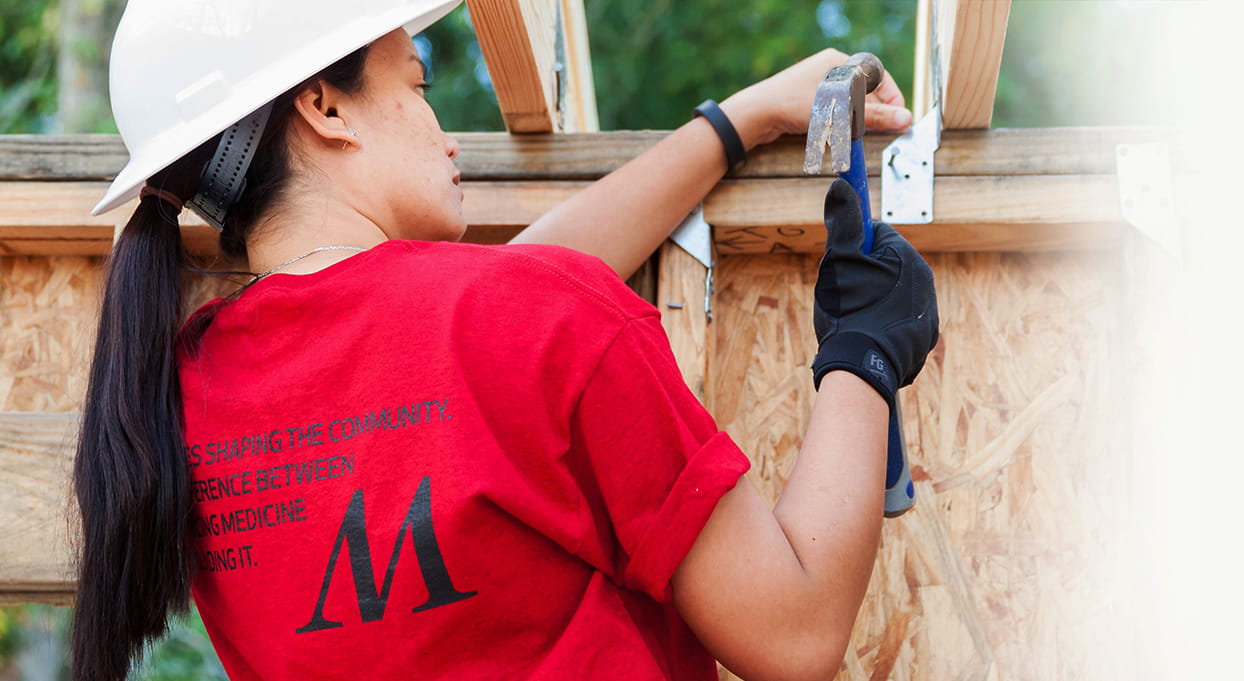

Community Benefits
Community Benefits focuses on providing access to high-quality, affordable health care services to indigent populations across Greater Houston while ensuring health equity for racial, ethnic and social minorities.




